PyOBO 0.9.2 Documentation
pyobo Package
A python package for handling and generating OBO.
Functions
|
Download a file if it doesn't exist. |
|
Get the OBO graph from a path. |
|
Get all of the terms from a OBO graph. |
|
Get alternative id to primary id mapping. |
|
Get all of the ancestors (parents) of the term as CURIEs. |
|
Get the definition for an entity. |
|
Get all of the descendants (children) of the term as CURIEs. |
|
Get equivalent CURIEs. |
|
Extract a single property for each term. |
|
Extract a single property for each term as a dictionary. |
|
Extract multiple properties for each term as a dictionary. |
|
Get all of the given relation. |
|
Get xrefs to a given target. |
|
Get hierarchy of parents as a directed graph. |
|
Get a mapping of descriptions. |
|
Get the OBO file and output a synonym dictionary. |
|
Get an identifier to name mapping for the OBO file. |
|
Get an identifier to species mapping. |
|
Get the OBO file and output a synonym dictionary. |
|
Get alternate identifiers. |
|
Get the set of identifiers for this prefix. |
|
Get the name for an entity. |
|
Get the name for a CURIE, if possible. |
|
Get a name to identifier mapping for the OBO file. |
|
Get the OBO for a given graph. |
|
Get the primary curie for an entity. |
|
Get the primary identifier for an entity. |
|
Get the priority CURIE mapped to the best namespace. |
|
Extract a set of properties for the given entity. |
|
Extract properties. |
|
Extract a single property for the given entity. |
|
Get the target identifier corresponding to the given relationship from the source prefix/identifier pair. |
|
Get relations from identifiers in the source prefix to target prefix with the given relation. |
|
Get all relations from the OBO. |
|
Get the species. |
|
Get xrefs from a source as an SSSOM dataframe. |
|
Get the subhierarchy for a given node. |
|
Get the synonyms for an entity. |
|
Get an identifier to name mapping for the typedefs in an OBO file. |
|
Get the PyOBO version string, including a git hash. |
|
Get the xref with the new prefix if a direct path exists. |
|
Get xrefs to a given target. |
|
Get all xrefs. |
|
Normalize a string given the prefix's labels and synonyms. |
|
Check that the first identifier has the second as an ancestor. |
|
Check if there's a plugin for converting the prefix. |
|
Check if there's a plugin for converting the prefix. |
|
Check that the first identifier has the second as a descendent. |
Get all modules in the PyOBO sources. |
|
|
Get all modules in the PyOBO sources. |
|
Parse a string that looks like a CURIE. |
Get parse results from an OBO graph. |
|
|
Get a converted PyOBO source. |
|
Get a converted PyOBO source. |
Classes
|
Wraps a graph and priority list to allow getting the best identifier. |
|
An OBO document. |
|
A utility for normalizing by names. |
|
A namespace, identifier, and label. |
|
A synonym with optional specificity and references. |
|
A type definition for synonyms in OBO. |
|
A term in OBO. |
|
A type definition in OBO. |
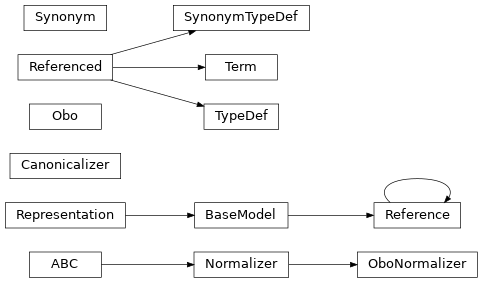
Class Inheritance Diagram